|
0 Comments
We are looking for a postdoc with experience in synthetic chemistry to work on several projects to develop inhibitors of chromatin remodeling. See the attached information below.
Postdoctoral Fellow in Chromatin Biology/Biochemistry/Pharmacology Department of Medicinal Chemistry and Molecular Pharmacology Purdue University Center for Cancer Research The Department of Medicinal Chemistry and Molecular Pharmacology Department at Purdue University Center for Cancer Research is seeking applicants for a full time postdoctoral fellow position in the laboratory of Prof. Emily Dykhuizen, who runs a highly interdisciplinary lab investigating and targeting chromatin remodelers. The postdoctoral fellow will be primarily developing and synthesizing chemical tools and inhibitors. If desired, potential projects can also include assay development, small molecule screening, medicinal chemistry, and chemical biology studies with small molecule candidates. Candidates must have a Ph.D in chemistry, biochemistry, medicinal chemistry or a closely related field. Candidates with experience in organic synthetic chemistry, medicinal chemistry, or chemical biology will be considered. This position is eligible for individual and dependent benefits, which includes medical, dental, disability, and life insurance. Please submit application, including a cover letter, CV and contact information (name, email, phone number) for at least 3 references to: Emily Dykhuizen, Ph.D. Associate Professor Department of Medicinal Chemistry and Molecular Pharmacology Purdue University HANS 105 201 S. University St. West Lafayette, IN 47907 [email protected] Dykhuizenlab.com Congrats to Sijie on successful defense of his dissertation, Good luck in your postdoc, Sijie. You will be missed!
Brayden Strohmier won the NIH Institutional NRSA Training Grant (T32) - Purdue Drug Discovery Training Program!
Congratulations! Thanks Amy for your devotion and help in the past years. You helped the lab so much, especially in these difficult times.
We're wishing you the best of luck in pharmacy school and in your future career! Welcome to MCMP graduate student Brayden Paul Strohmier! He will be working on the cellular effects of inhibiting BRD9 in prostate cancer cells and how those effects may change the resistance patterns of the prostate cancer to chemotherapeutics.
Congratulations, Aktan!
Aktan leaves for postdoc to cold spring harbor laboratory with Dr. Chris Vakoc. We will miss you! PBRM1 will be the death of me. It is a beautiful but poorly behaved protein. We've spent years trying to pin down reliable phenotypes in renal cancer that relate to specific biochemical function and have been humbled again and again. We've found dozens of cell lines with no detectable PBRM1 knockout phenotype. Then we've defined a phenotype in one cell line only to find a completely different phenotype in a very similar cell line. Then to make things worse, we've identified a phenotype in a cell line, only to find that phenotype lost or completely reversed when we take the knockout cell line back out of storage.
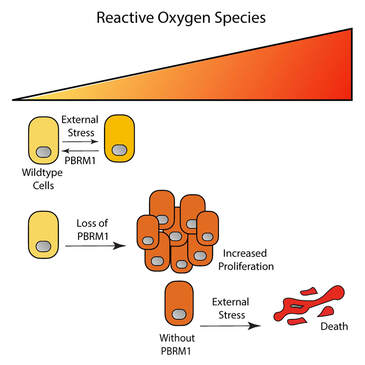
While we all know by now that chromatin regulators have context-dependent function, we really need to pin down this context in order to figure out how PBRM1 acts as a tumor suppressor in renal cancer (and other cancers). So in this paper we went back to untransformed cells to define PBRM1's role in maintaining epithelial cell maintenance. To make a long story short (if you want the long version, it is all in the paper) we find that PBRM1 maintains the expression of stress response genes in epithelial cells. Under low stress conditions loss of PBRM1 is growth-promoting, but under high stress conditions PBRM1 is cytoprotective and is required for the induction of stress response genes, the reduction of reactive oxygen species, and, eventually the initiation of apoptotic pathways. Now that we've found this, we understand why the same cell line responds differently to PBRM1 loss depending on the cell culture conditions, and why cell lines with inherently high internal stress (oxidative stress or oncogene addition) are so sensitive to PBRM1 knockdown. It's been a frustrating process, but a fruitful one (I think). Now we can start to piece together how PBRM1 responds to environmental signals to change gene expression. We have a lot of work to do, but I am hopeful that we have model that can explain all the varied and contradictory phenotypes reported for PBRM1. "PBRM1 Regulates Stress Response in Epithelial Cells." https://www.ncbi.nlm.nih.gov/pubmed/31077944 Libby is hooded and leaves the lab (sniff!). Libby is starting a postdoc in Ben Garcia's lab at UPenn and we are so happy for her! Congratulations!
Welcome to PULSe graduate student Surbhi Sood! She will be working on combining structural biology and genomics to decipher the connection between chromatin binding and transcriptional function of SWI/SNF chromatin remodelers. We are excited to get started!
|
AuthorDykhuizen Lab News Archives
May 2023
Categories |
Dykhuizen Lab
News and Pictures
Artwork by Libby Porter





 RSS Feed
RSS Feed